400 Dowman Dr.
Math and Science Center
Room: N246
Phone: 404-727-4930
Email: lfinzi@emory.edu
Transcription and related molecular motors
Elongation through transcription factors
Understanding how RNA polymerases transcribe through transcriptional factors that decorate DNA without interfering with their physiological role is paramount to our global understanding of maintenance of transcription regulation.
RNA polymerase I
RNA polymerase I (Pol I) is the eukaryotic polymerase that transcribes the genes that encode ribosomal RNA and is responsible for more than 60% of the cellular transcriptional activity. In collaboration with David Schneider (Univ. of Alabama, Birmingham), we are characterizing its elongation kinetics.
Topoisomerases
Topoisomerases are crucial in managing the level of DNA supercoiling largely arising from transcriptional activity of RNA polymerases. We study Type II topoisomerases in collaboration with David Dunlap using magnetic tweezers and labeling individual DNA molecules with super-paramagnetic beads.
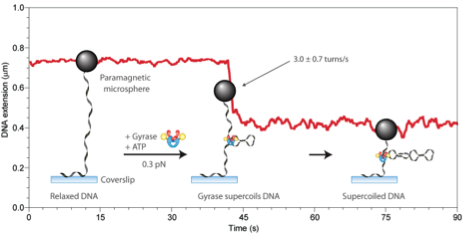
The experimental trace in the figure reveals the shortening of a DNA tether upon insertion of negative supercoils by Gyrase. The cartoons explain the the different parts of the trace.
Relevant Publications
| Authors | Title | Journal | Volume | Pages | Year |
| M.Fernandez, Q. Shao, C.Fountain, L. Finzi, and D. Dunlap | "E. coli gyrase fails to negatively supercoil diaminopurine-substituted DNA" | JMB | 2015 | ||
| Suleyman Ucuncuoglu, David A. Schneider, Eric Weeks, David Dunlap, and Laura Finzi | "Multiplexed, tethered particle microscopy for studies of DNA-enzyme dynamics" | Methods in Enzymology | accepted | ||
| S. Ucuncuoglu, K. L. Engel, P. K. Purohit, D. D. Dunlap, D. A. Schneider, L. Finzi | “Direct characterization of transcription elongation by RNA polymerase I” | Plos One | 2016 |
Relevant Techniques
| Method | Used for |
| AFM | to visualize transcriptional elongation |
| TPM | to monitor the rate of transcriptional elongation and the activity of topoisomerases |
| MTs | to study the activity of topoisomerases |
| PIFE | to monitor the dynamics of transcriptional elongation |